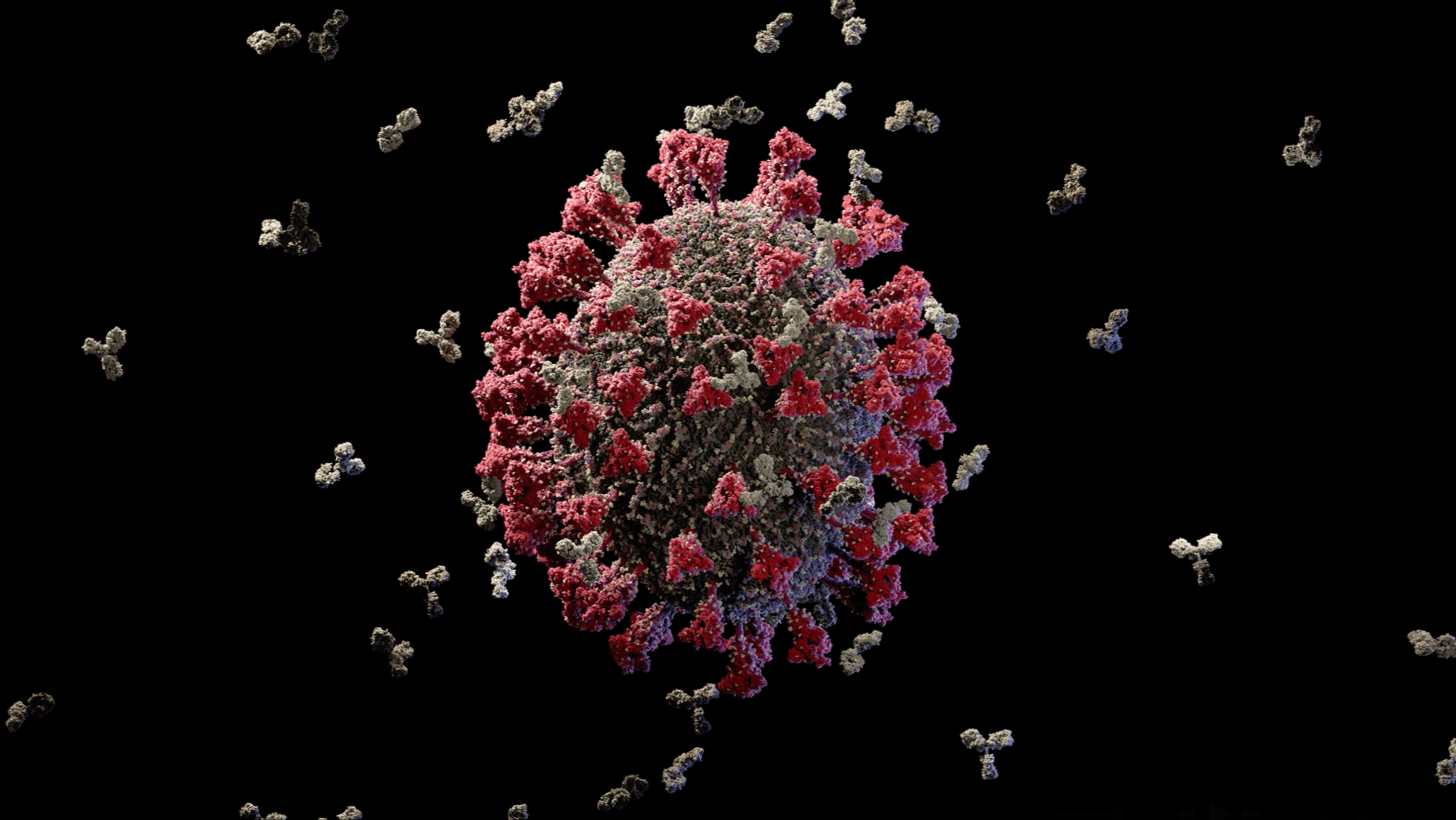Influenza A/H1N1
Virus 3D Model
Influenza A/H1N1
Virus 3D Model
We produced the first comprehensive and scientifically accurate 3D model of the Influenza A/H1N1 virus, commonly known as the seasonal flu, in collaboration with virologists from the National Center for Biotechnology in Madrid. The model reflects the state of scientific knowledge at the time of production and is based on extensive research in molecular biology, electron microscopy, and X-ray crystallography. The resulting visualization synthesizes fragmented experimental data into a coherent and biologically grounded representation of the viral particle.
We produced the first comprehensive and scientifically accurate 3D model of the Influenza A/H1N1 virus, commonly known as the seasonal flu, in collaboration with virologists from the National Center for Biotechnology in Madrid. The model reflects the state of scientific knowledge at the time of production and is based on extensive research in molecular biology, electron microscopy, and X-ray crystallography. The resulting visualization synthesizes fragmented experimental data into a coherent and biologically grounded representation of the viral particle.
Project Goal
Project Goal
Project Goal
Project Goal
The goal of this project was to create a scientifically accurate 3D visualization of the Influenza A/H1N1 virus that could clearly communicate its structural organization and molecular composition. The model was designed to support scientific education and public understanding by transforming complex and abstract virological data into an accessible visual format.
See full animation.
The goal of this project was to create a scientifically accurate 3D visualization of the Influenza A/H1N1 virus that could clearly communicate its structural organization and molecular composition. The model was designed to support scientific education and public understanding by transforming complex and abstract virological data into an accessible visual format.
See full animation.
The goal of this project was to create a scientifically accurate 3D visualization of the Influenza A/H1N1 virus that could clearly communicate its structural organization and molecular composition. The model was designed to support scientific education and public understanding by transforming complex and abstract virological data into an accessible visual format.
See full animation.
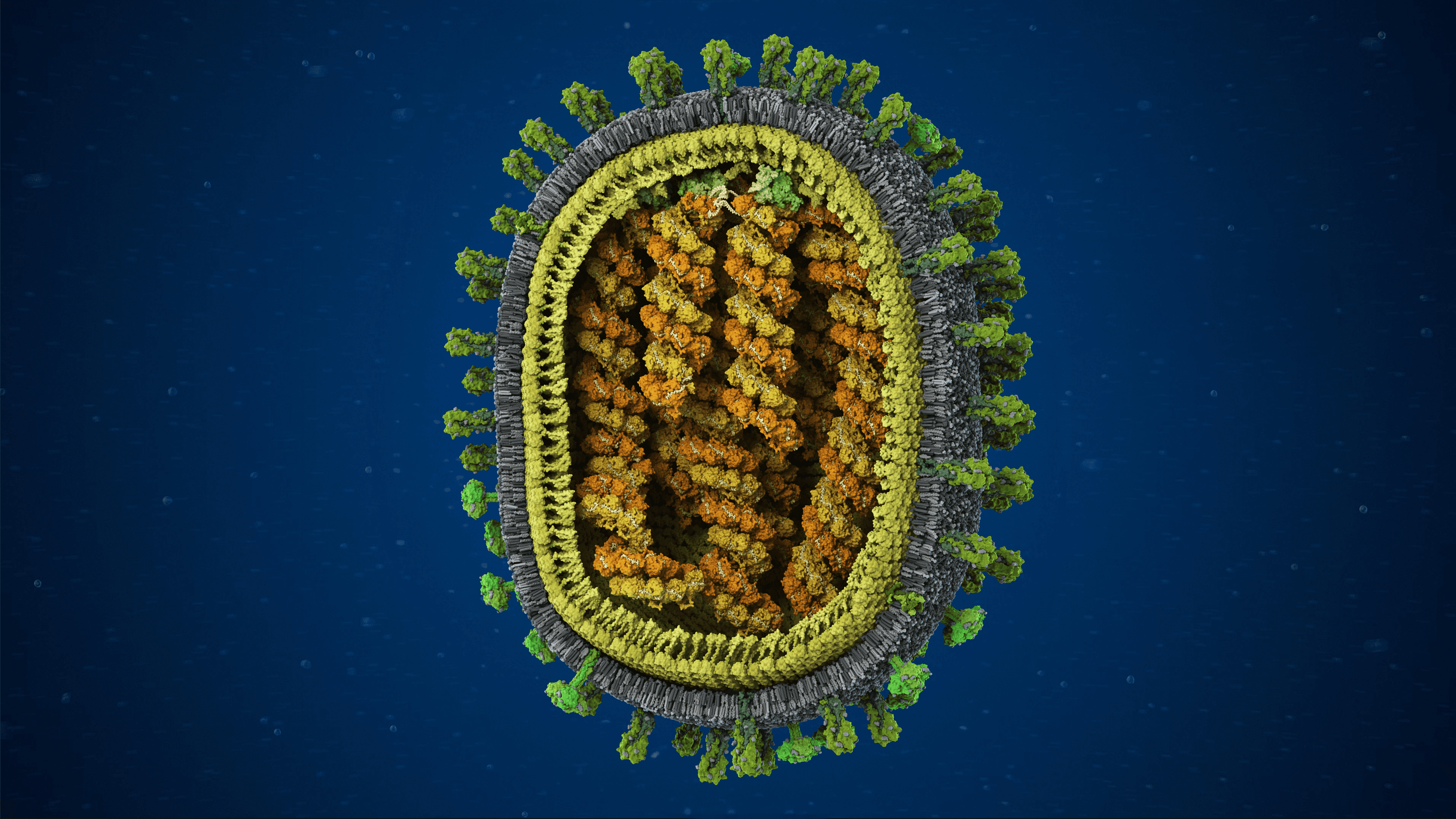
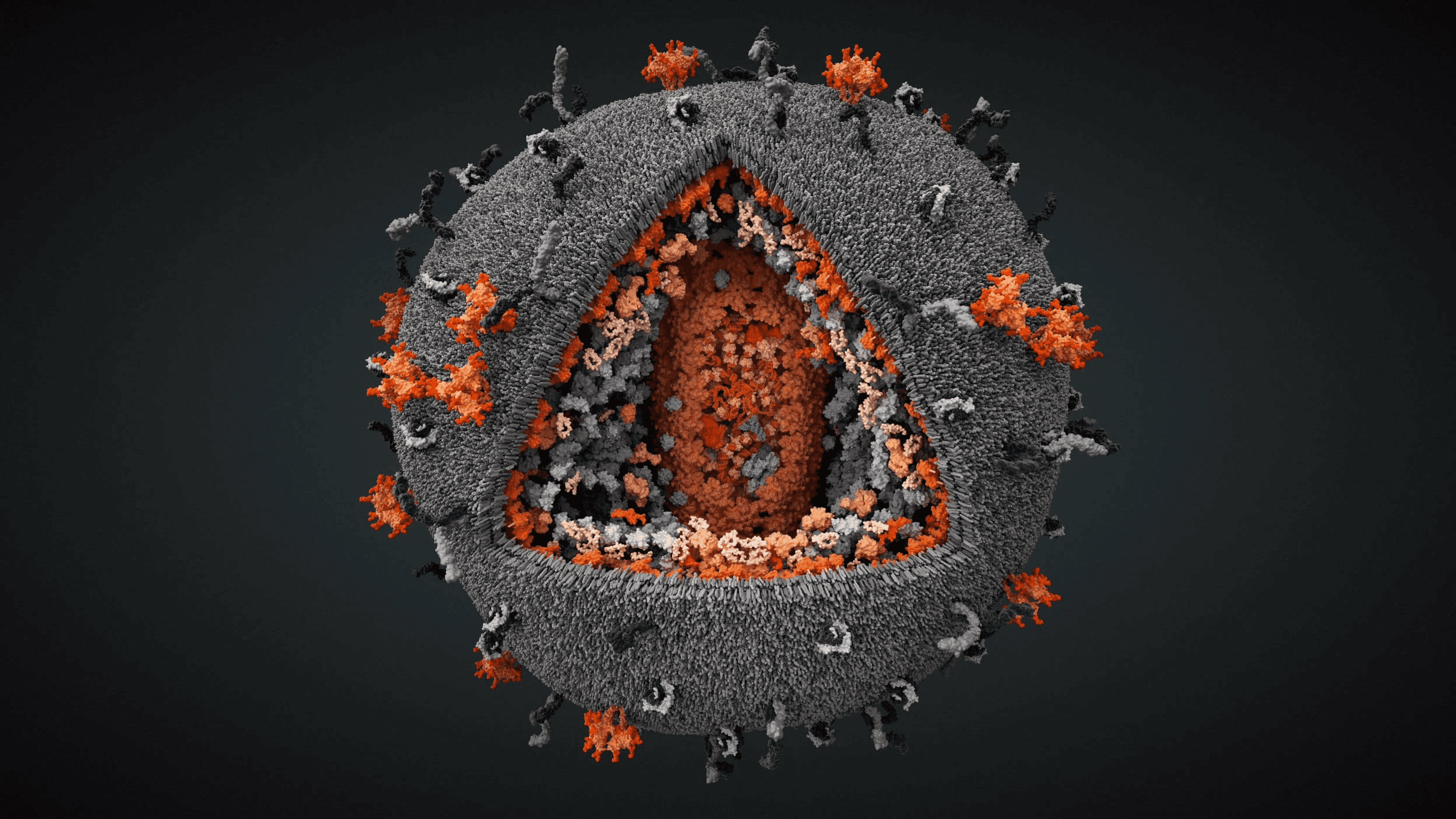
Influenza A/H1N1 virus with exposed core
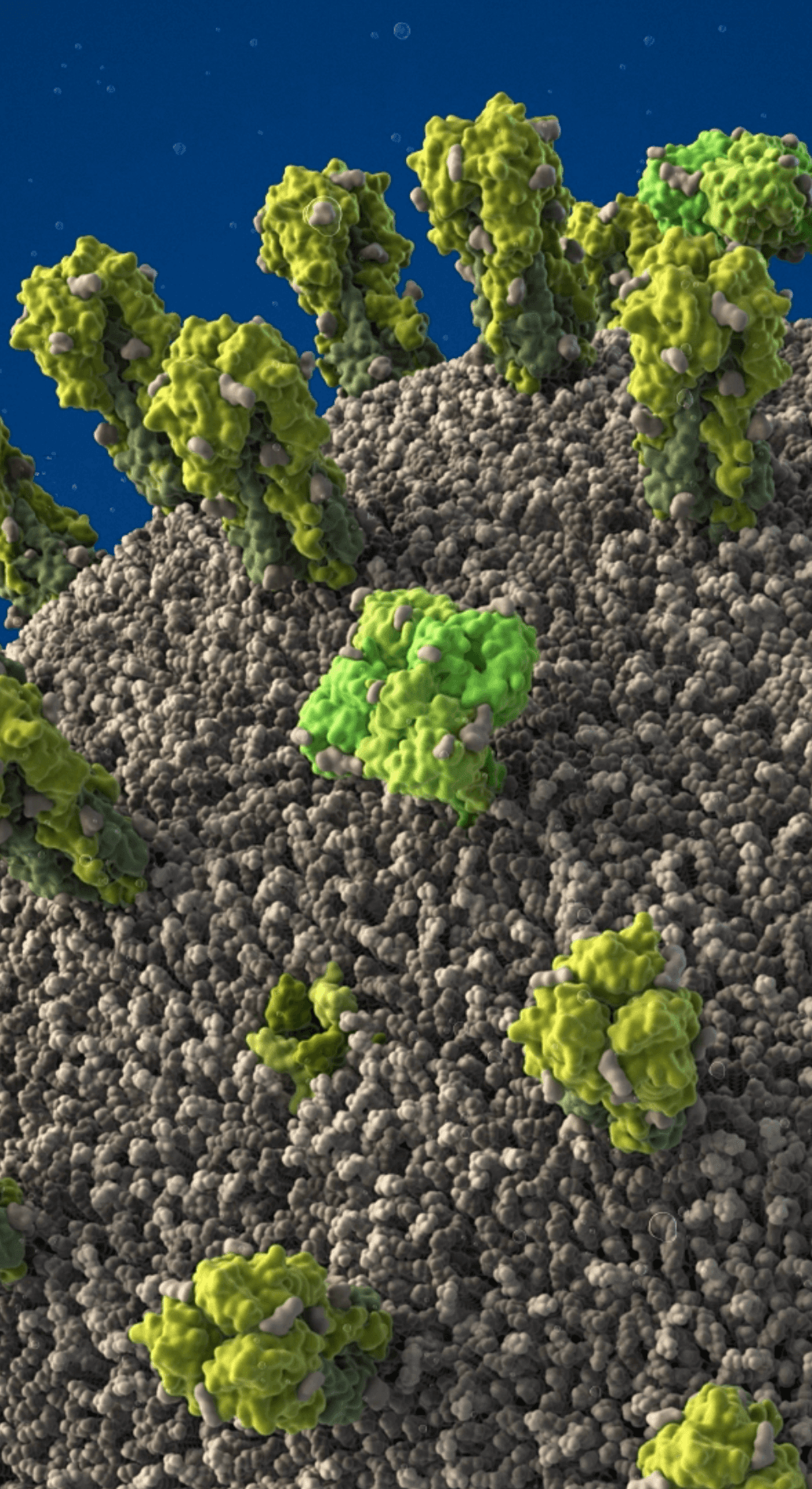
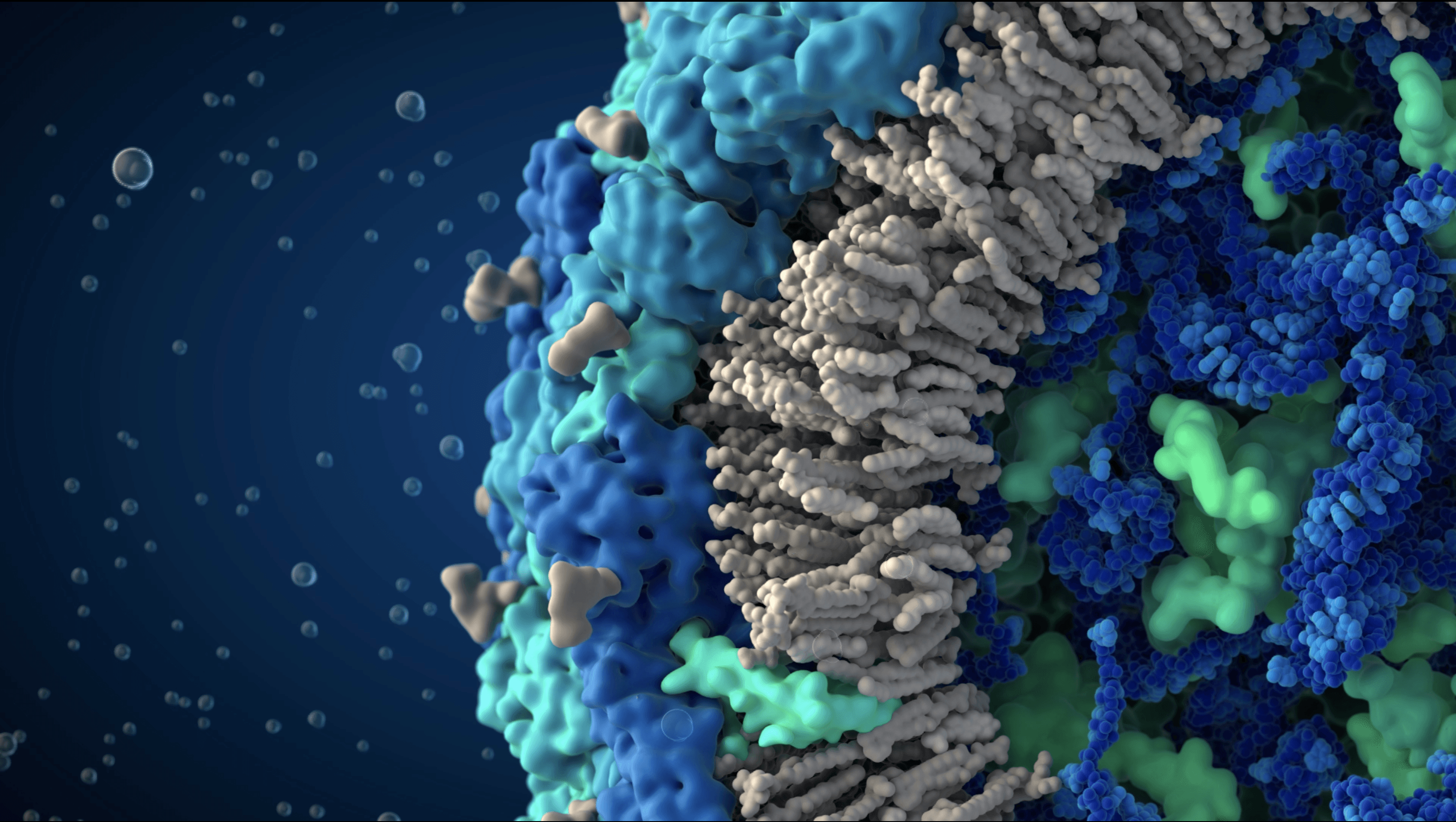
Proteins embedded in the viral envelope
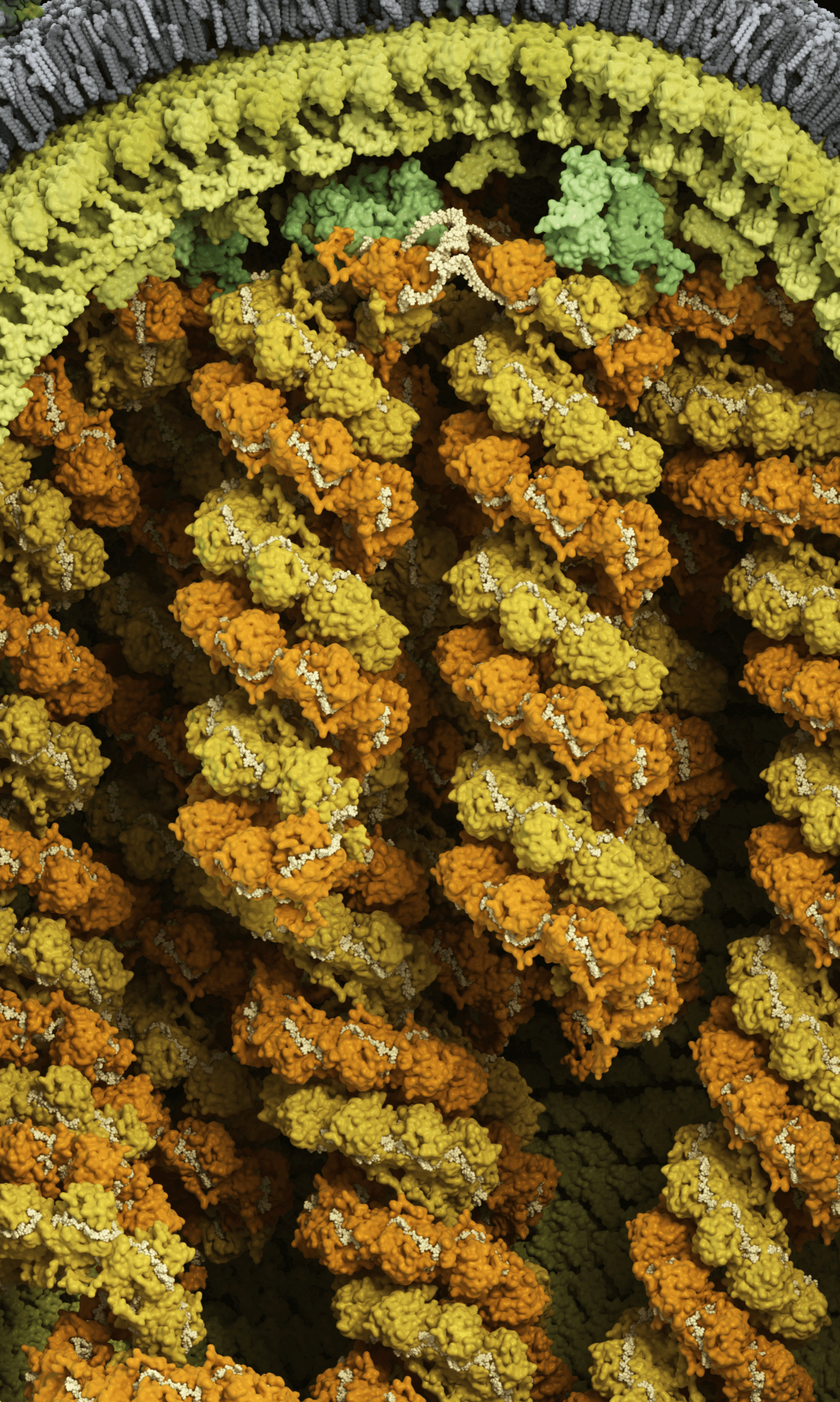
Genome surrounded by matrix and membrane
Genome surrounded by matrix and membrane
“ This is the most detailed 3D model we have of the influenza virus. Built by Visual Science that creates hyper-accurate 3D replicas of microscopic objects, the model aims to boost understanding of the bug.”
“ This is the most detailed 3D model we have of the influenza virus. Built by Visual Science that creates hyper-accurate 3D replicas of microscopic objects, the model aims to boost understanding of the bug.”
“ This is the most detailed 3D model we have of the influenza virus. Built by Visual Science that creates hyper-accurate 3D replicas of microscopic objects, the model aims to boost understanding of the bug.”
“ This is the most detailed 3D model we have of the influenza virus. Built by Visual Science that creates hyper-accurate 3D replicas of microscopic objects, the model aims to boost understanding of the bug.”

The Process
The Process
Animation type:
Animation type:
High-End 3D
High-End 3D
Project timline:
Project timline:
5 weeks
5 weeks
Find out which of our 26 scientific animation options works best for investor relations and communications:
The Influenza A/H1N1 model was developed through close collaboration between Visual Science and expert virologists to ensure accuracy and consistency with current research. Structural and molecular data were gathered from a wide range of peer-reviewed scientific studies, including findings from microscopy, crystallography, and molecular biology. Due to the absence of a complete experimental image of the virus, data from multiple sources were carefully combined to reconstruct the viral architecture. Molecular modeling techniques were used to assemble individual viral components into a unified 3D structure, which was then refined and animated to accurately convey spatial relationships and biological organization within the viral particle.
The Influenza A/H1N1 model was developed through close collaboration between Visual Science and expert virologists to ensure accuracy and consistency with current research. Structural and molecular data were gathered from a wide range of peer-reviewed scientific studies, including findings from microscopy, crystallography, and molecular biology. Due to the absence of a complete experimental image of the virus, data from multiple sources were carefully combined to reconstruct the viral architecture. Molecular modeling techniques were used to assemble individual viral components into a unified 3D structure, which was then refined and animated to accurately convey spatial relationships and biological organization within the viral particle.
Why did we use this
animation type?
Why did we use this
animation type?
The complex and microscopic nature of the Influenza A/H1N1 virus required a high-quality 3D visualization approach to accurately represent its structure. High-end 3D animation enabled precise control over geometry, lighting, materials, and camera movement, allowing the viral particle to be explored from multiple perspectives while preserving scientific integrity. This approach made it possible to present a clear and engaging scientific narrative that remains faithful to the underlying data and is suitable for both educational and research-focused audiences.
The complex and microscopic nature of the Influenza A/H1N1 virus required a high-quality 3D visualization approach to accurately represent its structure. High-end 3D animation enabled precise control over geometry, lighting, materials, and camera movement, allowing the viral particle to be explored from multiple perspectives while preserving scientific integrity. This approach made it possible to present a clear and engaging scientific narrative that remains faithful to the underlying data and is suitable for both educational and research-focused audiences.
Outcome:
Outcome:
The completed Influenza A/H1N1 model became a detailed and credible visual resource for understanding the structure of a ubiquitous human virus. Produced as part of Visual Science’s non-commercial Viral Park initiative, the project joined a growing collection of high-fidelity virus models, including HIV, Zika, and Sars-CoV-2 Virus. The influenza model demonstrates the effectiveness of rigorous 3D molecular visualization as a tool for communicating complex scientific knowledge and contributes to broader public and scientific engagement with virology.
The completed Influenza A/H1N1 model became a detailed and credible visual resource for understanding the structure of a ubiquitous human virus. Produced as part of Visual Science’s non-commercial Viral Park initiative, the project joined a growing collection of high-fidelity virus models, including HIV, Zika, and Sars-CoV-2 Virus. The influenza model demonstrates the effectiveness of rigorous 3D molecular visualization as a tool for communicating complex scientific knowledge and contributes to broader public and scientific engagement with virology.
Full animation
Contact us
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
Contact us
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
10/10
“
Fantastic team to partner with throughout the entire process. Excellent output and management.

Global Marketing Director, Pfizer
10/10
“
We are very pleased with Visual Science — they are a very responsive group to work with and the final product is exactly what we had envisioned.”

CEO, Atsena Therapeutics
10/10
“
The scientific expertise really showed in the discussions and final products. Everybody was responsive and great to work with, and the animations were both engaging and accurate!”

Program Manager, American Chemical Society
10/10
“
Visual Science team has just the right blend of scientific acumen and digital animation expertise to pull off these challenging projects.”

CEO, Apton Biosystems
9/10
“
Visual Science is a fantastic partner, capable of rendering the most complex science in compelling ways. They understood the science, and their production was excellent.”

VP, Head of Communications, Scorpion Tx
7/10
“
The structural biology models from Visual Science are stunning! Visual Science implemented our guidelines into beautiful scientific imagery for our website and scientific slide decks.”

Science Content Lead, Dewpoint Tx
9/10
“
I was very impressed with scientific knowledge of the Visual Science team, which led to a fantastic visualization of the MOA of our drug. Attention to scientific details was extraordinary.”

Vice President, R&D, Unicycive Tx
10/10
“
We were impressed by the scientific sophistication of the Visual Science team and their skill in capturing the concepts we wanted to communicate. The final video was exceptional.”

CSO, Enterin
10/10
“
The quality of the work from Visual Science was outstanding. They provide a bespoke service that was responsive to our requests and I would recommend them highly.”

CEO, UNITY Biotechnology
10/10
“
Beautiful work that is detailed and thoughtful. Great final output.”

Sr. Comms & IR Associate, Candel Tx
10/10
“
We were impressed by the Visual Science team’s ability to use exact protein structures to depict complex mechanisms in <90 seconds. The visuals are beautiful and tell a clear story.”

President & CEO, ROME Tx
10/10
“
Excellent scientific partner and first class imagery. This team runs a tight ship and the process is clear. The tech and final product are first rate.”

VP, Corp Dev, Revolution Medicines
Contact us
Contact us
Visual Science is an award-winning medical animation and digital scientific communications company, trusted by leading biotech and pharmaceutical organizations since 2007, including J&J, Pfizer, Novartis, Roche, Takeda, Gilead, AbbVie, and 100+ others.
We specialize in science-grade MoA and MoD videos, medical animation, and scientific storytelling, as well as digital and AI-driven solutions for Medical Affairs, marketing, corporate communications, and investor relations.
© 2026, Visual Science. All rights reserved.
Visual Science is an award-winning medical animation and digital scientific communications company, trusted by leading biotech and pharmaceutical organizations since 2007, including J&J, Pfizer, Novartis, Roche, Takeda, Gilead, AbbVie, and 100+ others.
We specialize in science-grade MoA and MoD videos, medical animation, and scientific storytelling, as well as digital and AI-driven solutions for Medical Affairs, marketing, corporate communications, and investor relations.
© 2026, Visual Science. All rights reserved.
Visual Science is an award-winning medical animation and digital scientific communications company, trusted by leading biotech and pharmaceutical organizations since 2007, including J&J, Pfizer, Novartis, Roche, Takeda, Gilead, AbbVie, and 100+ others.
We specialize in science-grade MoA and MoD videos, medical animation, and scientific storytelling, as well as digital and AI-driven solutions for Medical Affairs, marketing, corporate communications, and investor relations.
© 2026, Visual Science. All rights reserved.
Visual Science is an award-winning medical animation and digital scientific communications company, trusted by leading biotech and pharmaceutical organizations since 2007, including J&J, Pfizer, Novartis, Roche, Takeda, Gilead, AbbVie, and 100+ others.
We specialize in science-grade MoA and MoD videos, medical animation, and scientific storytelling, as well as digital and AI-driven solutions for Medical Affairs, marketing, corporate communications, and investor relations.
© 2026, Visual Science. All rights reserved.